I want to create a matrix that displays the r.squared coefficient of determination of some predictions made over the years and the actual values.
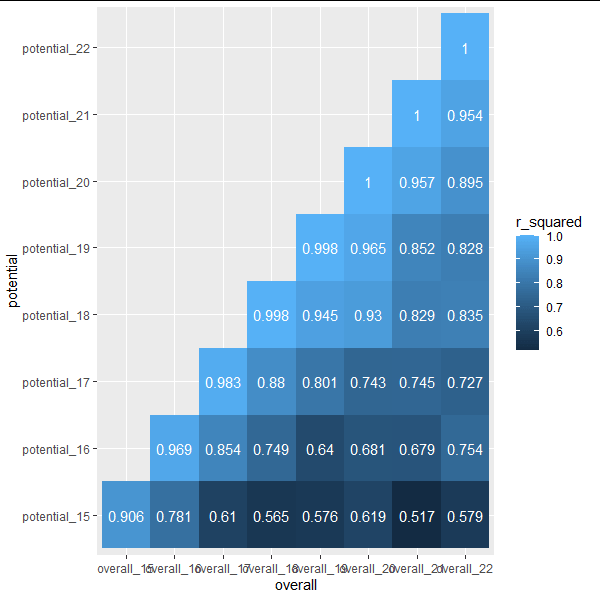
My goal is to display a matrix that looks something like this.

The only way I found is to make multiple lists, calculate each row/ column individually using map2_dbl(l.predicted_line1, l.actual, ~ summary(lm(.x ~ .y))$r.squared), and then add the resulting vectors in a matrix with some code. This would create 9 lists, which I want to avoid.
Is there any way of doing this in a more efficiently?
#sample data
l.actual <- list(
overall_15 = c(59,65,73,73,64,69,64,69,63,NA,82,60,NA,73,NA,73,73,NA,69,
69,71,66,65,70,72,72,NA,64,69,67,64,71,NA,62,62,71,67,63,64,76,72),
overall_16 = c(60,68,75,74,68,71,NA,72,64,69,82,66,64,77,NA,71,72,NA,69,
69,75,67,71,73,73,73,NA,66,NA,69,65,70,76,NA,67,71,72,64,65,76,73),
overall_17 = c(63,68,NA,74,72,72,NA,73,66,69,83,67,64,76,NA,71,73,NA,70,
70,79,NA,73,72,NA,NA,NA,NA,NA,70,NA,70,77,NA,68,74,74,66,64,75,69),
overall_18 = c(NA,68,NA,78,73,72,NA,72,68,67,86,NA,62,75,65,71,71,67,71,
71,76,NA,71,71,NA,NA,74,NA,71,NA,NA,68,74,NA,67,75,74,65,NA,72,NA),
overall_19 = c(NA,NA,NA,77,73,72,NA,71,69,66,87,63,62,73,65,NA,NA,NA,NA,
NA,75,NA,NA,67,NA,NA,73,NA,NA,NA,NA,NA,74,NA,NA,74,74,65,NA,68,NA),
overall_20 = c(NA,NA,NA,77,NA,NA,NA,72,71,66,87,NA,NA,NA,65,NA,NA,NA,70,
70,75,NA,NA,NA,NA,NA,NA,NA,NA,NA,NA,NA,74,NA,66,71,73,NA,NA,69,NA),
overall_21 = c(NA,67,NA,76,NA,69,NA,73,69,65,85,NA,NA,NA,NA,NA,NA,NA,NA,
NA,75,NA,NA,NA,NA,NA,69,NA,NA,NA,NA,NA,73,NA,67,68,72,NA,NA,68,NA),
overall_22 = c(NA,NA,NA,75,NA,NA,NA,75,67,65,84,NA,NA,NA,NA,NA,NA,NA,68,
68,73,NA,NA,NA,NA,NA,NA,NA,NA,NA,NA,NA,NA,NA,67,69,71,NA,NA,68,NA)
)
l.predicted <- list(
potential_15 = c(59,68,74,76,65,75,64,72,66,NA,85,60,NA,76,NA,73,75,NA,71,
71,71,67,65,70,72,72,NA,68,74,67,64,71,NA,62,62,71,71,63,67,78,72),
potential_16 = c(60,71,75,75,68,73,NA,74,66,69,83,66,64,77,NA,71,74,NA,70,
70,76,67,71,73,73,73,NA,66,NA,69,65,70,76,NA,67,71,72,64,66,76,73),
potential_17 = c(63,69,NA,75,72,72,NA,73,69,69,83,67,64,76,NA,71,73,NA,70,
70,79,NA,73,72,NA,NA,NA,NA,NA,70,NA,70,77,NA,68,74,74,66,64,75,69),
potential_18 = c(NA,68,NA,78,73,72,NA,72,69,67,86,NA,62,75,65,71,71,67,71,
71,76,NA,71,71,NA,NA,74,NA,71,NA,NA,68,74,NA,67,75,74,65,NA,72,NA),
potential_19 = c(NA,NA,NA,77,73,72,NA,71,70,66,87,63,62,73,65,NA,NA,NA,NA,
NA,75,NA,NA,67,NA,NA,73,NA,NA,NA,NA,NA,74,NA,NA,74,74,65,NA,68,NA),
potential_20 = c(NA,NA,NA,77,NA,NA,NA,72,71,66,87,NA,NA,NA,65,NA,NA,NA,70,
70,75,NA,NA,NA,NA,NA,NA,NA,NA,NA,NA,NA,74,NA,66,71,73,NA,NA,69,NA),
potential_21 = c(NA,67,NA,76,NA,69,NA,73,69,65,85,NA,NA,NA,NA,NA,NA,NA,NA,
NA,75,NA,NA,NA,NA,NA,69,NA,NA,NA,NA,NA,73,NA,67,68,72,NA,NA,68,NA),
potential_22 = c(NA,NA,NA,75,NA,NA,NA,75,67,65,84,NA,NA,NA,NA,NA,NA,NA,68,
68,73,NA,NA,NA,NA,NA,NA,NA,NA,NA,NA,NA,NA,NA,67,69,71,NA,NA,68,NA)
)
>Solution :
Here is a solution using some tidyverse packages. The key thing is to use the function expand_grid() to get all combinations of the elements of each list. This results in a tibble with two named list columns. Next we can use mutate() to pull out the names of the list and assign them to new columns, and extract the numeric IDs. Use filter() to retain only the rows where potential is less than or equal to overall. Finally get the R-squared for each row using your suggested code, and plot. (Note I did not try too hard to get the plot to look just like yours.)
library(purrr)
library(dplyr)
library(ggplot2)
library(tidyr)
r_squared_combinations <- expand_grid(l.actual, l.predicted) %>%
mutate(overall = names(l.actual),
potential = names(l.predicted),
overall_n = as.numeric(gsub('overall_', '', overall)),
potential_n = as.numeric(gsub('potential_', '', potential))) %>%
filter(potential_n <= overall_n) %>%
mutate(r_squared = map2_dbl(l.predicted, l.actual, ~ summary(lm(.x ~ .y))$r.squared))
ggplot(r_squared_combinations, aes(x = overall, y = potential, fill = r_squared, label = round(r_squared, 3))) +
geom_tile() +
geom_text(color = 'white')
Side note: incidentally the base function expand.grid() would work about as well as tidyr::expand_grid() but expand_grid() returns a tibble by default which may be more convenient if you are using tidyverse functions otherwise.